These functions extend the functionality of dplyr::sample_n() and
dplyr::slice_sample() by allowing for repeated sampling of data.
This operation is especially helpful while creating sampling
distributions—see the examples below!
Usage
rep_sample_n(tbl, size, replace = FALSE, reps = 1, prob = NULL)
rep_slice_sample(
.data,
n = NULL,
prop = NULL,
replace = FALSE,
weight_by = NULL,
reps = 1
)Arguments
- tbl, .data
Data frame of population from which to sample.
- size, n, prop
sizeandnrefer to the sample size of each sample. Thesizeargument torep_sample_n()is required, while inrep_slice_sample()sample size defaults to 1 if not specified.prop, an argument torep_slice_sample(), refers to the proportion of rows to sample in each sample, and is rounded down in the case thatprop * nrow(.data)is not an integer. When usingrep_slice_sample(), please only supply one ofnorprop.- replace
Should samples be taken with replacement?
- reps
Number of samples to take.
- prob, weight_by
A vector of sampling weights for each of the rows in
.data—must have length equal tonrow(.data). Forweight_by, this may also be an unquoted column name in.data.
Value
A tibble of size reps * n rows corresponding to reps
samples of size n from .data, grouped by replicate.
Details
rep_sample_n() and rep_slice_sample() are designed to behave similar to
their dplyr counterparts. As such, they have at least the following
differences:
In case
replace = FALSEhavingsizebigger than number of data rows inrep_sample_n()will give an error. Inrep_slice_sample()having suchnorprop > 1will give warning and output sample size will be set to number of rows in data.
Note that the dplyr::sample_n() function has been superseded by
dplyr::slice_sample().
Examples
library(dplyr)
#>
#> Attaching package: ‘dplyr’
#> The following objects are masked from ‘package:stats’:
#>
#> filter, lag
#> The following objects are masked from ‘package:base’:
#>
#> intersect, setdiff, setequal, union
library(ggplot2)
library(tibble)
# take 1000 samples of size n = 50, without replacement
slices <- gss |>
rep_slice_sample(n = 50, reps = 1000)
slices
#> # A tibble: 50,000 × 12
#> # Groups: replicate [1,000]
#> replicate year age sex college partyid hompop hours income
#> <int> <dbl> <dbl> <fct> <fct> <fct> <dbl> <dbl> <ord>
#> 1 1 1994 49 female no degree ind 4 40 $25000 …
#> 2 1 1985 51 female no degree dem 4 28 $25000 …
#> 3 1 2010 51 female degree rep 4 60 $25000 …
#> 4 1 2016 58 female no degree rep 3 22 $25000 …
#> 5 1 2010 60 male degree ind 2 45 $25000 …
#> 6 1 2004 48 male no degree rep 4 89 $25000 …
#> 7 1 2000 68 male no degree dem 2 60 $25000 …
#> 8 1 1998 58 female no degree ind 4 24 $10000 …
#> 9 1 1996 49 female degree dem 3 60 $25000 …
#> 10 1 1989 29 female degree dem 4 30 $25000 …
#> # ℹ 49,990 more rows
#> # ℹ 3 more variables: class <fct>, finrela <fct>, weight <dbl>
# compute the proportion of respondents with a college
# degree in each replicate
p_hats <- slices |>
group_by(replicate) |>
summarize(prop_college = mean(college == "degree"))
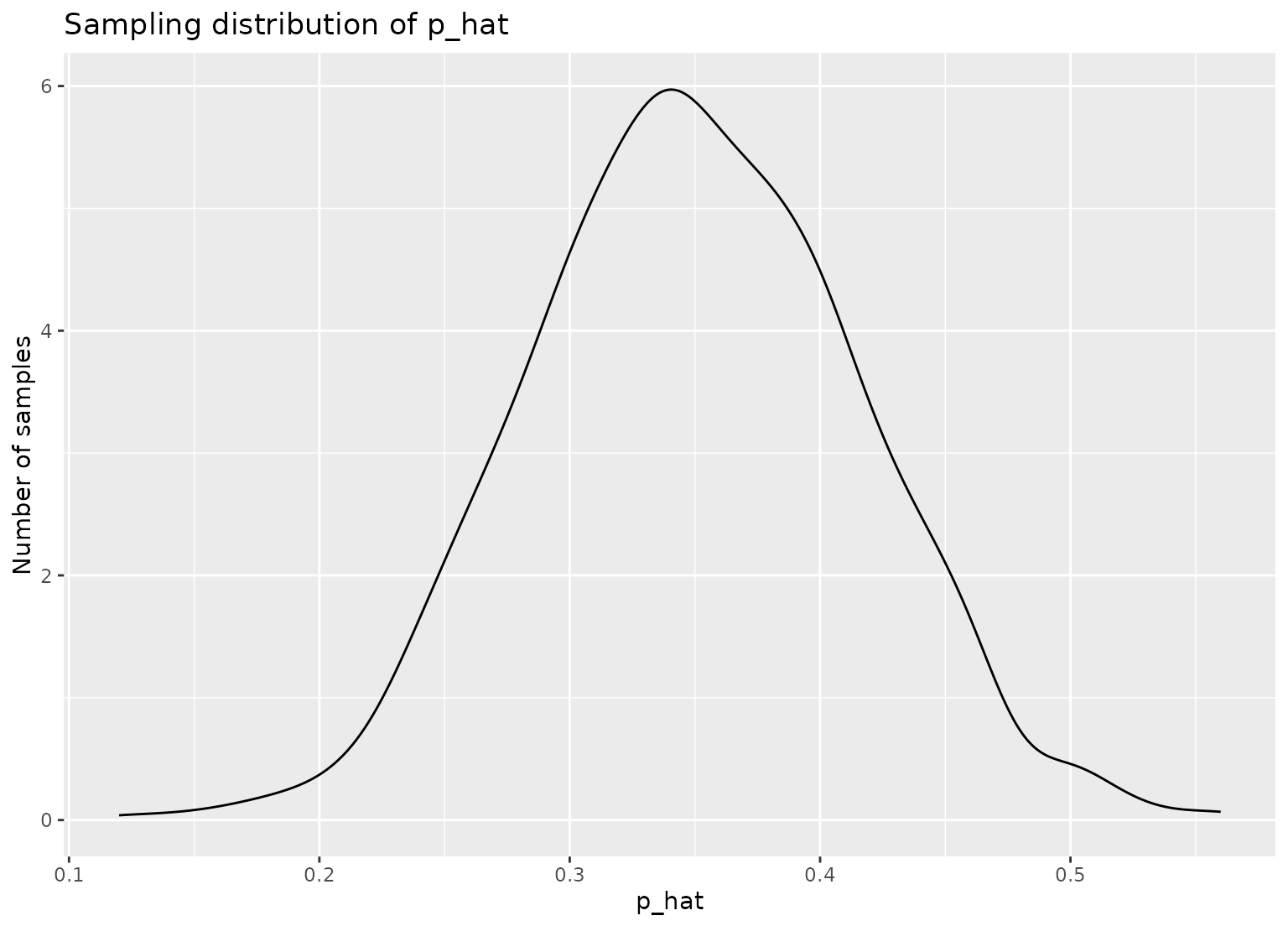
# plot sampling distribution
ggplot(p_hats, aes(x = prop_college)) +
geom_density() +
labs(
x = "p_hat", y = "Number of samples",
title = "Sampling distribution of p_hat"
)
 # sampling with probability weights. Note probabilities are automatically
# renormalized to sum to 1
df <- tibble(
id = 1:5,
letter = factor(c("a", "b", "c", "d", "e"))
)
rep_slice_sample(df, n = 2, reps = 5, weight_by = c(.5, .4, .3, .2, .1))
#> # A tibble: 10 × 3
#> # Groups: replicate [5]
#> replicate id letter
#> <int> <int> <fct>
#> 1 1 5 e
#> 2 1 2 b
#> 3 2 1 a
#> 4 2 5 e
#> 5 3 1 a
#> 6 3 3 c
#> 7 4 1 a
#> 8 4 3 c
#> 9 5 1 a
#> 10 5 2 b
# alternatively, pass an unquoted column name in `.data` as `weight_by`
df <- df |> mutate(wts = c(.5, .4, .3, .2, .1))
rep_slice_sample(df, n = 2, reps = 5, weight_by = wts)
#> # A tibble: 10 × 4
#> # Groups: replicate [5]
#> replicate id letter wts
#> <int> <int> <fct> <dbl>
#> 1 1 2 b 0.4
#> 2 1 5 e 0.1
#> 3 2 4 d 0.2
#> 4 2 2 b 0.4
#> 5 3 3 c 0.3
#> 6 3 5 e 0.1
#> 7 4 3 c 0.3
#> 8 4 1 a 0.5
#> 9 5 5 e 0.1
#> 10 5 3 c 0.3
# sampling with probability weights. Note probabilities are automatically
# renormalized to sum to 1
df <- tibble(
id = 1:5,
letter = factor(c("a", "b", "c", "d", "e"))
)
rep_slice_sample(df, n = 2, reps = 5, weight_by = c(.5, .4, .3, .2, .1))
#> # A tibble: 10 × 3
#> # Groups: replicate [5]
#> replicate id letter
#> <int> <int> <fct>
#> 1 1 5 e
#> 2 1 2 b
#> 3 2 1 a
#> 4 2 5 e
#> 5 3 1 a
#> 6 3 3 c
#> 7 4 1 a
#> 8 4 3 c
#> 9 5 1 a
#> 10 5 2 b
# alternatively, pass an unquoted column name in `.data` as `weight_by`
df <- df |> mutate(wts = c(.5, .4, .3, .2, .1))
rep_slice_sample(df, n = 2, reps = 5, weight_by = wts)
#> # A tibble: 10 × 4
#> # Groups: replicate [5]
#> replicate id letter wts
#> <int> <int> <fct> <dbl>
#> 1 1 2 b 0.4
#> 2 1 5 e 0.1
#> 3 2 4 d 0.2
#> 4 2 2 b 0.4
#> 5 3 3 c 0.3
#> 6 3 5 e 0.1
#> 7 4 3 c 0.3
#> 8 4 1 a 0.5
#> 9 5 5 e 0.1
#> 10 5 3 c 0.3
